|
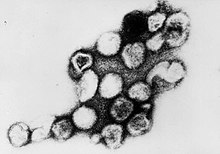
Rubella virus
Rubella virus (RuV) is the pathogenic agent of the disease rubella, transmitted only between humans via the respiratory route, and is the main cause of congenital rubella syndrome when infection occurs during the first weeks of pregnancy. Rubella virus, scientific name Rubivirus rubellae, is a member of the genus Rubivirus and belongs to the family of Matonaviridae, whose members commonly have a genome of single-stranded RNA of positive polarity which is enclosed by an icosahedral capsid. As of 1999[update] the molecular basis for the causation of congenital rubella syndrome was not yet completely clear, but in vitro studies with cell lines showed that rubella virus has an apoptotic effect on certain cell types. There is evidence for a p53-dependent mechanism.[2] TaxonomyRubella virus (Rubivirus rubellae) is assigned to the Rubivirus genus.[1] Matonaviridae familyUntil 2018, Rubiviruses were classified as part of the family Togaviridae, but have since been changed to be the sole genus of the family Matonaviridae. This family is named after George de Maton, who in 1814 first distinguished rubella from measles and scarlet fever.[3] The change was made by the International Committee on the Taxonomy of Viruses (ICTV), the central governing body for viral classification. Matonaviridae remains part of the realm that it was already in as Togaviridae, Riboviria, because of its RNA genome and RNA dependent RNA polymerase.[3] Other rubivirusesIn 2020, Ruhugu virus and Rustrela virus joined Rubella virus as second and third of only three members of the genus Rubivirus.[4] Neither of them are known to infect people.[5] MorphologyWhile alphavirus virions are spherical and contain an icosahedral nucleocapsid, RuV virions are pleiomorphic and do not contain icosahedral nucleocapsids.[3] PhylogenyICTV analyzed the sequence of RuV and compared its phylogeny to that of togaviruses. They concluded:
StructureThe spherical virus particles (virions) of Matonaviridae have a diameter of 50 to 70 nm and are covered by a lipid membrane (viral envelope), derived from the host cell membrane. There are prominent "spikes" (projections) of 6 nm composed of the viral envelope proteins E1 and E2 embedded in the membrane.[6] The E1 glycoprotein is considered immunodominant in the humoral response induced against the structural proteins and contains both neutralizing and hemagglutinating determinants.
Capsid proteinInside the lipid envelope is a capsid of 40 nm in diameter. The capsid protein (CP) has different functions.[8] Its main tasks are the formation of homooligomeres to form the capsid, and the binding of the genomic RNA. Further is it responsible for the aggregation of RNA in the capsid, it interacts with the membrane proteins E1 and E2 and binds the human host-protein p32 which is important for replication of the virus in the host.[9] As opposed to alphaviruses the capsid does not undergo autoproteolysis, rather is it cut off from the rest of the polyprotein by the signal-peptidase. Production of the capsid happens at the surface of intracellular membranes simultaneously with the budding of the virus.[10] GenomeThe positive-strand RNA genome has 9,762 nucleotides and encodes 2 nonstructural polypeptides (p150 and p90) within its 5′-terminal two-thirds and 3 structural polypeptides (C, E2, and E1) within its 3′-terminal one-third.[11] Both envelope proteins E1 and E2 are glycosylated. There are three sites that are highly conserved in Matonaviruses: a stem-and-loop structure at the 5' end of the genome, a 51-nucleotide conserved sequence near the 5' end of the genome and a 20-nucleotide conserved sequence at the subgenomic RNA start site. Homologous sequences are present in the rubella genome.[11] The genome encodes several non-coding RNA structures; among them is the rubella virus 3' cis-acting element, which contains multiple stem-loops, one of which has been found to be essential for viral replication.[12] The only significant region of homology between rubella and the alphaviruses is located at the NH2 terminus of non structural protein 3. This sequence has helicase and replicase activity. In the rubella genome these occur in the opposite orientation to that found in the alphaviruses indicating that a genome rearrangement has occurred. The genome has the highest G+C content of any currently known single stranded RNA virus (~70%).[13] Despite this high GC content its codon use is similar to that of its human host. ReplicationThe viruses attach to the cell surface via specific receptors and are taken up by an endosome being formed. At the neutral pH outside of the cell the E2 envelope protein covers the E1 protein. The dropping pH inside the endosome frees the outer domain of E1 and causes the fusion of the viral envelope with the endosomal membrane. Thus, the capsid reaches the cytosol, decays and releases the genome The +ssRNA (positive, single-stranded RNA) at first only acts as a template for the translation of the non-structural proteins, which are synthesized as a large polyprotein and are then cut into single proteins. The sequences for the structural proteins are first replicated by the viral RNA polymerase (Replicase) via a complementary −ssRNA as a template and translated as a separate short mRNA. This short subgenomic RNA is additionally packed in a virion.[14] Translation of the structural proteins produces a large polypeptide (110 kDa). This is then endoproteolytically cut into E1, E2 and the capsid protein. E1 and E2 are type I transmembrane proteins which are transported into the endoplasmatic reticulum (ER) with the help of an N-terminal signal sequence. From the ER the heterodimeric E1·E2-complex reaches the Golgi apparatus, where the budding of new virions occurs (unlike alpha viruses, where budding occurs at the plasma membrane. The capsid proteins on the other hand stay in the cytoplasm and interact with the genomic RNA, together forming the capsid.[15]
TransmissionRuV is transmitted via respiration between humans.[3] EpidemiologyOn the basis of differences in the sequence of the E1 protein, two genotypes have been described which differ by 8 - 10%. These have been subdivided into 13 recognised genotypes - 1a, 1B, 1C, 1D, 1E, 1F, 1G, 1h, 1i, 1j, 2A, 2B and 2C. For typing, the WHO recommends a minimum window that includes nucleotides 8731 to 9469.[16] Genotypes 1a, 1E, 1F, 2A and 2B have been isolated in China. Genotype 1j has only been isolated from Japan and the Philippines. Genotype 1E is found in Africa, the Americas, Asia and Europe. Genotype 1G has been isolated in Belarus, Côte d'Ivoire and Uganda. Genotype 1C is endemic only in Central and South America. Genotype 2B has been isolated in South Africa. Genotype 2C has been isolated in Russia. Literature
References
External links |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||